Advanced Pandas: Optimize speed and memory
Nowadays the Python data analysis library Pandas is widely used across the world. It started mostly as a data exploration and experimentation tool but is slowly transitioning to be used in a production-like setting. It is for instance used in scheduled data extraction and model training via Airflow, or even in a streaming fashion to prepare the data for model inference. We could argue Pandas or Python are not suitable tools for production environments. Nevertheless it is useful to know how to get the most out of it.
Pandas is a very powerful tool, but needs mastering to gain optimal performance. In this post you will learn how to optimize processing speed and memory usage with the following outline:
- Index Optimization
- Vectorize Operations
- Memory Optimization
- Filter Optimization
Throughout this article we will make use of Kaggle’s public Berlin Airbnb dataset, which consist of house listings, neighbourhoods and reviews. Let’s look at a couple of rows and columns from the listings set:
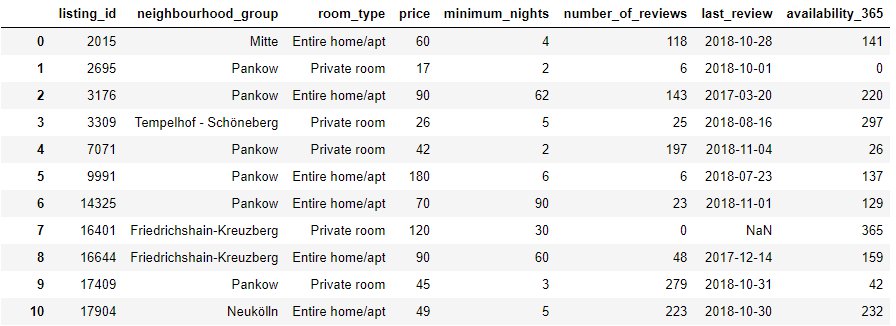
1. Index Optimalization
Pandas has optimized operations based on indices, allowing for faster lookup or merging tables based on indices. In the following example we merge the reviews table with the listings table, first using a column to merge on, then using the index.
Even when having to set the index, merging on indices is faster. Let’s see the differences when looking up a value.
Single lookups using indices outperform other methods with great margin. When retrieving a single value, using .at[] is faster than using .loc[].
2. Vectorize Operations
Vectorization is the process of executing operations on entire arrays. Similarly to numpy, Pandas has built in optimizations for vectorized operations. It is advised to avoid for loops when working with dataframes, since read and write operations are costly. However it is not always possible to vectorize, so we will also show what the best iterative options for pandas are by comparing .iloc[], .iterrows(), .loc[]and.map()/.apply().
In the following example we want to normalize the price column between 0 and 1, using the different operations stated above.
The difference in computation speed is enormous, with the vectorized solution being ~82000 times faster than using .iloc[].
While this is a very simple example, it matters most with these simple operations, where value lookup covers most of the computation time.
Writing vectorized code can be difficult for complex problems. Often intermediate solutions have to be stored in the dataframe, which can be memory inefficient. A good alternative for doing complex operations is taking the data out of the dataframe and loop through numpy arrays. Let’s look at an example where we use business rules to determine the most suitable Airbnb for us, first by looping using .loc[], then converting to numpy arrays and lastly by using some Pandas vectorization magic.
.loc[]
numpy
vectorized
We see that looping through numpy arrays is a fast alternative to vectorized operations, which is useful when vectorization becomes too complex. For a limited number of samples the computation time using numpy becomes less than having a vectorized approach. We sampled data from our listings dataframe and computed the average time of 50 runs for both approaches.
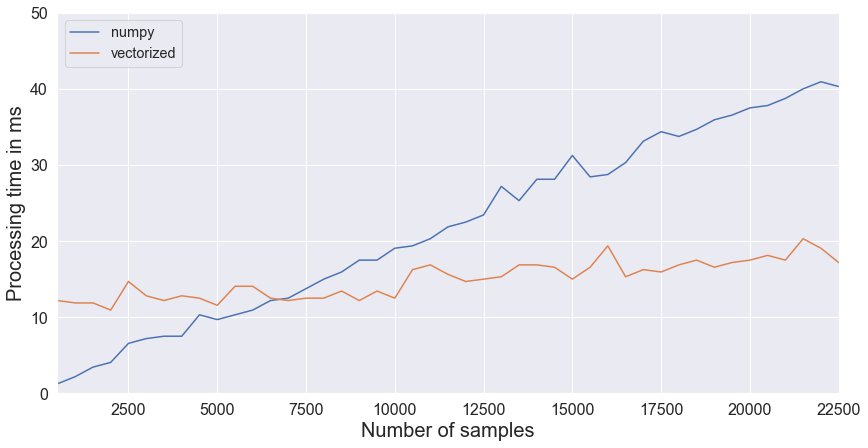
Both methods seem to have a linear time complexity, however numpy looping has a steeper slope, but a lower intercept. The difference in intercept is that with Pandas vectorization more new columns are created, which is costly.
3. Memory Optimization
One of the drawbacks of Pandas is that by default the memory consumption of a DataFrame is inefficient. When reading in a csv or json file the column types are inferred and are defaulted to the largest data type (int64, float64, object).
Our listings DataFrame is build via a csv, which has the following properties:
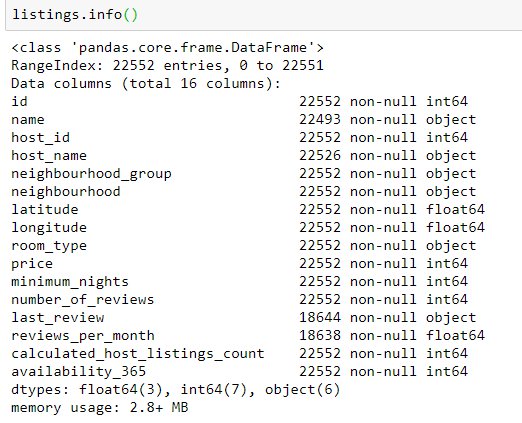
All columns have the previously stated largest data types. We can see some obvious improvements here, for instance the availability_365 has only 365 possible values, so it can be downcasted to an int16 (-32,768 to +32,767).
We defined some functions that will downcast the columns automatically to the smallest possible datatype without losing any information. For strings we make use of the pandas category column type if the amount of unique strings cover less than half the total amount of strings. This should be tailored to your own needs, since defining a category restricts adding unknown values to that column. Date columns we cast to the pandas datetime dtype. It does not reduce memory usage, but enables time based operations.
Now we can simply optimize our listings dataframe by callingoptimized_listings = optimize(listings, [‘last_review’])
This results in the following dataframe:
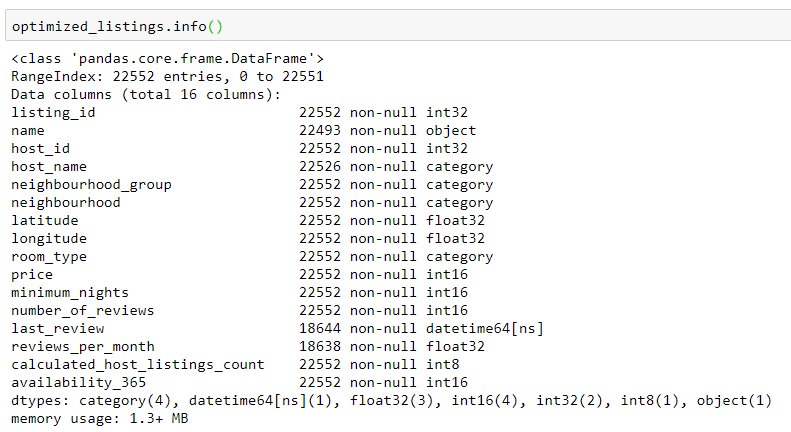
Only two columns have the same dtypes as before, so optimizing was definitely worth it. The total memory usage have dropped from 2.8MB to 1.3MB, improving over 50%. For this toy example it is unnecessary, but very helpful for large dataframes.
An alternative to the custom optimization code is specifying the dtypes when reading the csv. This requires knowledge about the value ranges per column.
To retain these new column types you can store the dataframe in a type which can hold column type information, such as pickle. This will also reduce the physical size of the file as can be seen in the example below.

Disk usage between optimized csv and pickle files
4. Filter Optimization
When chaining multiple operations it is worthwhile to think about which operations to execute first. Filter steps should be executed as early as possible.
Even when making inner joins between dataframes, it is worthwhile to filter before merging. If we sample our listings with .sample(frac=0.2) and then merge the reviews to it, we see that its more efficient to filter the reviews first.
If the discrepancy between the two dataframes is large this can save a lot of time.
Summary
In this post we’ve seen that small changes in our code can significantly speed up computations and reduce the memory usage of dataframes. Here are a few key takeaways:
- Use indexing as much as possible for merging and value lookup
- a) Use vectorized operations
b) When looping is unavoidable, use native numpy, or .map() for simple operations - a) Use the stated memory optimization code to greatly reduce memory
b) Store large dataframes as a pickle file to retain the column types and reduce disk usage - Always filter data in early stages


